Gene editing in BEAS-2B cell line facilitates respiratory researches.
Human bronchial epithelium cell line (BEAS-2B) was derived from normal bronchial epithelium obtained from the autopsy of non-cancerous individuals. This cell line comes from the ATCC CRL-9609, it is also known as bronchial epithelial cell line. The BEAS-2B cell line was initially infected by an adenovirus 12-SV40 hybrid virus, and the immortalized cell line of BEAS-2B established by continuous cell passage. BEAS-2B cell retains the ability to differentiate from serum response, this ability can be used to screen chemical or biological agents that induce or affect differentiation and carcinogenesis. Therefore, BEAS-2B cells are the ideal cell model for the study of drug metabolic activation and respiratory tumors and molecular mechanisms.
2.Applications of BEAS-2B cell line
2.1 The BEAS-2B cells shared a similar potential with hMSCs on both osteogenic and adipogenic differentiation
After culturing for 21 days under each differentiation induction condition, Oil Red O was used to stain lipid vacuoles in the differentiated adipocytes and Alizarin Red S was used to stain calcium deposits in the differentiated osteocytes. The result shows that both BEAS-2B and hMSC1 cells showed an almost equivalent positive staining for either osteogenesis or adipogenesis. These results indicate that, similar to hMSC1, BEAS-2B exhibited a strong capability of differentiating into osteocytes and adipocytes after each differentiation induction .
2.2 The BEAS-2B can be used to screen chemical and biological agents.
BEAS-2B has been extensively used to study cellular and molecular mechanisms involved in lung carcinogenesis, including the role of epithelial-mesenchymal transition (EMT) in lung carcinogenesis, as well as pneumococcal infections. In addition, the BEAS-2B cell line has been utilized as an in vitro cell model for assaying or screening various chemicals and biological agents with potential pulmonary toxicity or lung carcinogenicity.
3.CRISPR-U ™ mediated Gene editing in BEAS-2B cell line
Genome editing and the creation of cell models can advance researches in the area of functional genomics, signaling pathways, metabolism, cell death, drug discovery, drug response, and cancer research, etc. BEAS-2B is the ideal cell model for the study of drug metabolic activation and respiratory tumors and molecular mechanisms. So far, CRISPR/Cas9 technology has been used by many researchers to study cancer characteristics, disclosure of drug resistance mechanism, cancer treatment, cell death research, functional genomics, signaling pathway, drug discovery, drug response, and cell therapy, etc. CRISPR-U™ (based on CRISPR/Cas9 technology), developed by Ubigene, is more efficient than general CRISPR/Cas9 in double-strand breaking, and CRISPR-U™ can greatly improve the efficiency of homologous recombination, easily achieve gene editing in vitro and in vivo. It has great advantages for gene editing of BEAS-2B cells.
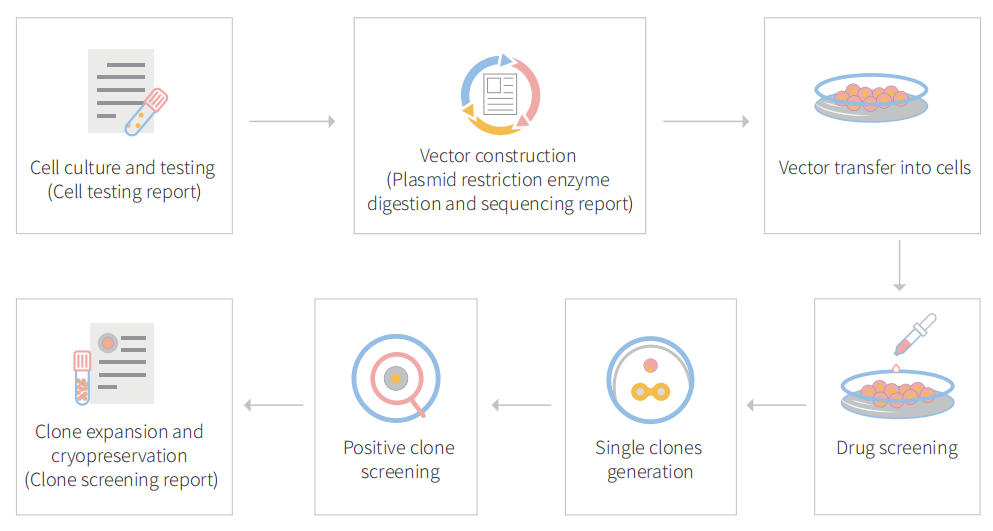
Figure 1 Gene editing in BEAS-2B cells with CRISPR-U™
3.1 Gene knockout in BEAS-2B cell line with CRISPR-U™
gRNA and Cas9 would be transferred into cells by virus transduction, liposome transfection, or nucleofection. After drug screening, single clones would be isolated. Positive clones would be validated by sequencing. Specific case is as shown below:
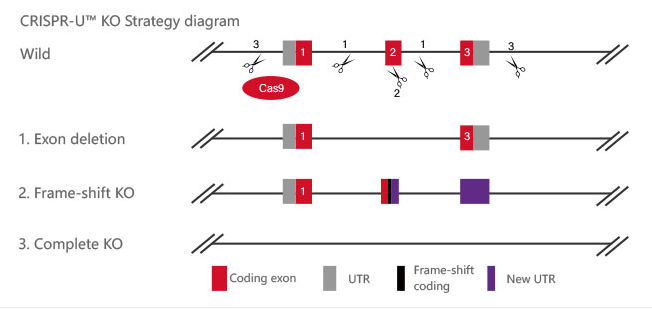
Figure 2 Knockout Strategies of BEAS-2B cell line
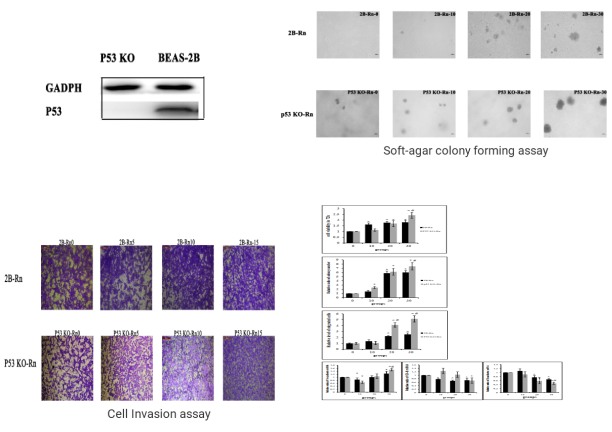
P53 gene is an important factor in the malignant transformation of BEAS-2B cells by radon. The P53 gene was knocked out in human normal lung epithelial cells (BEAS-2B) and radon was treated for gene knockout BEAS-2B cells and normal BEAS-2B cells, respectively. The researchers found earlier malignant transformation of BEAS-2B cells from radon after knocking the P53 gene. The BEAS-2B cell (P53 KO-Rn) proliferate faster (BEAS-2B <0.05), earlier cloning (P <0.05), more invasive (P <0.05), and earlier changes in mRNA expression levels of epithelial-interstitial transformation (EMT) -related genes (P <0.05). The results show that radon irradiation leads to malignant transformation of BEAS-2B cells and that P53 plays an important role in radon gas-induced lung cancer.
Reference :http://www.paper.edu.cn/releasepaper/content/201910-45
3.2 Gene KI/point mutation BEAS-2B cell line
BEAS-2B cells would be co-transfected with gRNA, Cas9 and donor vector by electroporation, and then screened. After drug screening, single clones would be generated. Positive clones would be validated by sequencing. Specific cases are as shown below:
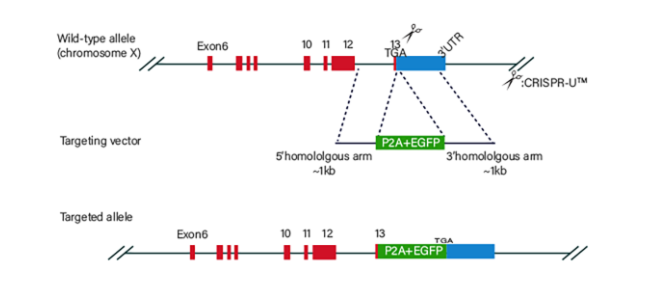
Figure 3 Knock-in Strategies of BEAS-2B cell line
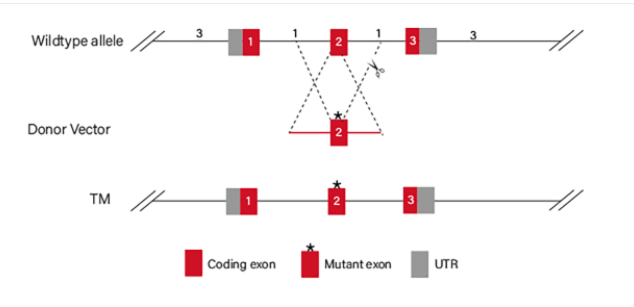
Figure 4 Point mutation Strategies of BEAS-2B cell line
(1)Case of point mutation cell line-1
To study the specific influence of NRASQ61K mutant on angiogenesis, NRASQ61K was knocked into normal (non-neoplastic) human bronchial epithelial cells (BEAS-2B). Compared to wild type (WT) cells, NRASQ61K expressing cells growing on CAM were characterized by a disorganized vasculature with several hemorrhagic areas. To confirm the proangiogenic effects of NRASQ61K in vivo, a Matrigel plug assay was performed in athymic nude mice. As shown in Figure 5B, NRASQ61K -expressing cells demonstrated a marked increase in angiogenesis in the plugs, as indicated by a bright red color. However, angiogenesis was negligible and hemoglobin levels were lower in the WT cells. As shown in Figure 5C, cells spontaneously form capillary-like structures on Matrigel, which is a critical step in tumor angiogenesis. Therefore, NRASQ61K expression has the effect on tubulogenesis.
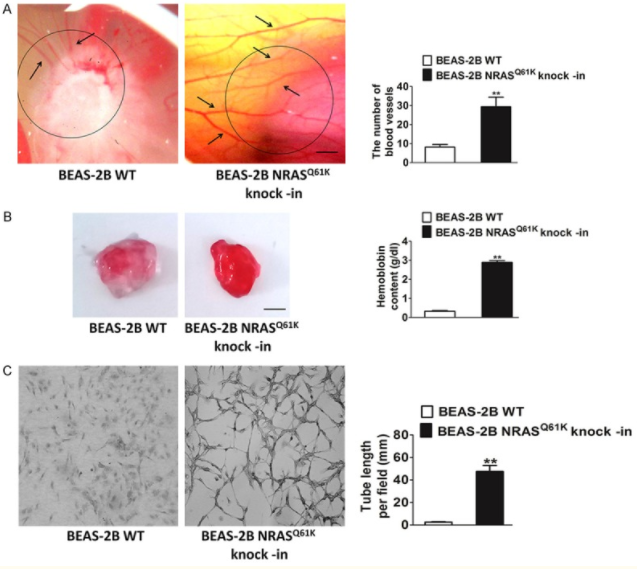
Figure 5 Knock-in of NRASQ61K into BEAS-2B
(2)Case of point mutation cell line-2
Arsenite Exposure Led to Activation of NFAT but Not AP-1 and NFκB in Beas-2B Cells, the COX-2 gene promoter region contains NFκB, AP-1, and NFAT binding sites, which can be recognized by these transcription factors and in turn lead to COX-2 transcription. To determine whether NFAT regulated COX-2 expression through direct binding to the COX-2 promoter region, point mutation of the two NFAT binding sites in the promoter region of COX-2-Luc reporter was carried out by CRISPR/Cas9. The results demonstrated that this mutation resulted in impairment of the COX-2 transcription induced by arsenite exposure. And the researchers found that it may be noted that NFAT activation by arsenite in the time course studies reached a peak earlier, as compared with that of COX-2 induction. Because the exposure of arsenite only causes NFAT activation in BEAS-2B cells, not AP-1 and NFκB activation. Thus, the activation of NFAT by arsenite is the reason for COX-2 induction in BEAS-2B cells.
In addition, to confirm the important role of NFAT3 in arsenite-induced COX-2 expression in BEAS-2B cells, the researchers built and used the siNFAT3. They used the cells for 10 μm NFAT selective inhibitors were pretreated and then exposed to 20μm arsenite. The results demonstrated that stable transfection of siNFAT3 in Beas-2B cells resulted in a dramatic reduction of NFAT3 protein expression in Beas-2B cells (Fig. 6b). Specific knockdown of NFAT3 expression by siNFAT3 blocked arsenite-induced COX-2 transcription and protein expression (Fig. 6, e and h).
These results distinctly demonstrate that NFAT3 is a major mediator for COX-2 induction by arsenite in Beas-2B cells.
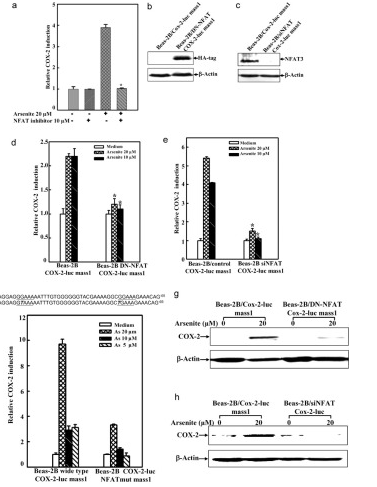
Figure 6 Point mutation cases of BEAS-2B cell line
References:
[1]Liu, Shuang, et al. "miR�106b�5p targeting SIX1 inhibits TGF�β1�induced pulmonary fibrosis and epithelial�mesenchymal transition in asthma through regulation of E2F1." International Journal of Molecular Medicine 47.3 (2021): 1-1.
[2]Sawada J, Li F, Komatsu M. R-Ras protein inhibits autophosphorylation of vascular endothelial growth factor receptor 2 in endothelial cells and suppresses receptor activation in tumor vasculature. J Biol Chem. 2015;290:8133–8145
[3]He, Jun, et al. "Arsenic-induced metabolic shift triggered by the loss of miR-199a-5p through Sp1-dependent DNA methylation." Toxicology and applied pharmacology 378 (2019): 114606.
[4]Pircher A, Hilbe W, Heidegger I, Drevs J, Tichelli A, Medinger M. Biomarkers in tumor angiogenesis and anti-angiogenic therapy. Int J Mol Sci. 2011;12:7077–7099.
[5]Song, Zhaowei, Fenghai Liu, and Jie Zhang. "Targeting NRASQ61K mutant delays tumor growth and angiogenesis in non-small cell lung cancer." American journal of cancer research 7.4 (2017): 831.
[6]W.R. Chappell, B.D. Beck, K.G. Brown, R. Chaney, R. Cothern, C.R. Cothern, K.J. Irgolic, D.W. North, I. Thornton, T.A. Tsongas. Environ. Health Perspect., 105 (1997), pp. 1060-1067
[7]Ding, Jin, et al. "Cyclooxygenase-2 induction by arsenite is through a nuclear factor of activated T-cell-dependent pathway and plays an antiapoptotic role in Beas-2B cells." Journal of Biological Chemistry 281.34 (2006): 24405-24413.
[8]H.Y. Chiou, Y.M. Hsueh, K.F. Liaw, S.F. Horng, M.H. Chiang, Y.S. Pu, J.S. Lin, C.H. Huang, C.J. Chen. Cancer Res., 55 (1995), pp. 1296-1300
[9]Van Den Heuvel, Rosette, et al. "Identification of PM10 characteristics involved in cellular responses in human bronchial epithelial cells (Beas-2B)." Environmental research 149 (2016): 48-56.












